This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
What are protein domains?
Protein domains are units of a protein that have specific structure and function [1]. Some genes can have only one domain, while others have many. Domains can be used to predict functions of a protein based on its similarities with other known proteins with similar/the same domains [2]. A few functions a protein domain might have are binding to DNA and interaction with other proteins. One way a protein domain can be studied is by comparing homologous proteins across species, as not all will be exactly the same. This helps to understand structure and function of proteins by examining their roles in the different organisms.
How to find domains:
To find domains databases such as Pfam and SMART are used.
How to find domains:
To find domains databases such as Pfam and SMART are used.
MITF Protein Domains
MITF domains from Pfam:
Pfam shows the MITF domain (green), HLH domain (red), and DFU3371 domain (blue).
Pfam shows the MITF domain (green), HLH domain (red), and DFU3371 domain (blue).
MITF domains from SMART:
SMART shows the only domain is HLH.
SMART shows the only domain is HLH.
|
MITF domain
This domain is found at the N-terminus of MITF. However, there is no further information known about this domain [3]. |
DUF3371 domain
At this time, this domain has an unknown function [3].
At this time, this domain has an unknown function [3].
Domain Analysis
Pfam and SMART both show different results for the number of domains that are part of the MITF protein. Pfam and SMART use different thresholds for what is considered to be a domain, therefore they do not always show the same results [3]. They both agree on the HLH domain being apart of the protein. There is not much known about the other two domains (MITF and DUF3371) so there may need to be more studies done before they would be considered domains in both Pfam and SMART.
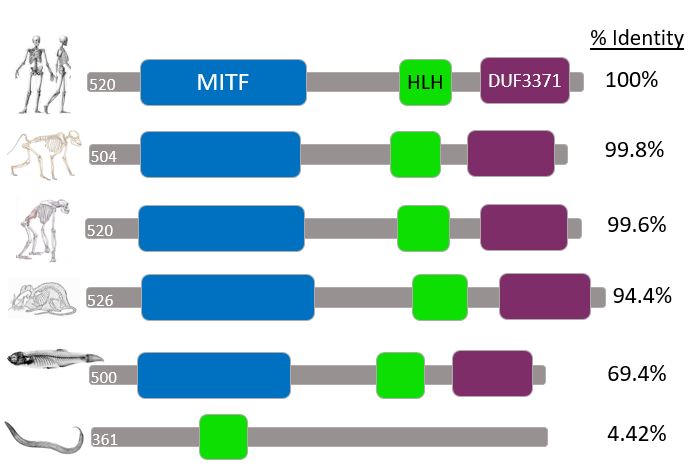
Domain comparisons across species
Comparison Analysis
MITF domains are highly conserved across these different species. Since the % Identity of the protein homologues is very high for mammals it is logical that their protein domains would also be highly conserved among the group. The notable exception here is C. elegans. However, since the % Identity is very low (4.42%) this too makes sense.
Sources for finding domains:
References
1. Protein Domains, Domain Assignment, domain identification, the CATH database. (n.d.). Retrieved February 25, 2017, from http://www.proteinstructures.com/Structure/Structure/protein-domains.html
2. Lee, D., Redfern, O., & Orengo, C. (2007). Predicting protein function from sequence and structure. Nature Reviews Molecular Cell Biology,8(12), 995-1005. doi:10.1038/nrm2281
3. Home page. (n.d.). Retrieved February 25, 2017, from http://pfam.xfam.org/
1. Protein Domains, Domain Assignment, domain identification, the CATH database. (n.d.). Retrieved February 25, 2017, from http://www.proteinstructures.com/Structure/Structure/protein-domains.html
2. Lee, D., Redfern, O., & Orengo, C. (2007). Predicting protein function from sequence and structure. Nature Reviews Molecular Cell Biology,8(12), 995-1005. doi:10.1038/nrm2281
3. Home page. (n.d.). Retrieved February 25, 2017, from http://pfam.xfam.org/