This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Introduction
Waardenburg syndrome type 2A (WS2A) is a rare autosomal dominant syndrome that affects the hearing and also leads to benign pigmentation abnormalities [1]. Waardenburg syndrome is caused by the loss of the MITF gene. MITF (microphthalmia-associated transcription factor) is located on chromosome 3 and has been associated with WS2A cases. MITF induces the conversion of fibroblasts into melanocytes, a mature melanin-forming cell, and is involved in the differentiation of these cells [2].
When mutated there is a loss of function of these genes and it results in reduced pigmentation, as seen in the hair and eyes of those affected by WS2A. It has been shown that the mutated MITF gene loses the ability to bind to the promoter, therefore the necessary genes for the melanocyte cell differentiation cannot be activated [2]. Mutations in MITF can often cause a patch of white hair and brilliant blue coloring of eyes, and sometimes only one eye is affected [1].
One interesting aspect of WS2A, is the fact that it is autosomal dominant. This means that if an individual has one copy of the gene mutated they will have the disease. However, in WS2A individuals they can have incomplete penetrance of pigmentation loss. This is seen by the pigmentation loss in only one eye of some individuals. The molecular mechanisms for this incompletely penetrant phenotype in the eyes is unknown.
When mutated there is a loss of function of these genes and it results in reduced pigmentation, as seen in the hair and eyes of those affected by WS2A. It has been shown that the mutated MITF gene loses the ability to bind to the promoter, therefore the necessary genes for the melanocyte cell differentiation cannot be activated [2]. Mutations in MITF can often cause a patch of white hair and brilliant blue coloring of eyes, and sometimes only one eye is affected [1].
One interesting aspect of WS2A, is the fact that it is autosomal dominant. This means that if an individual has one copy of the gene mutated they will have the disease. However, in WS2A individuals they can have incomplete penetrance of pigmentation loss. This is seen by the pigmentation loss in only one eye of some individuals. The molecular mechanisms for this incompletely penetrant phenotype in the eyes is unknown.
Aim 1
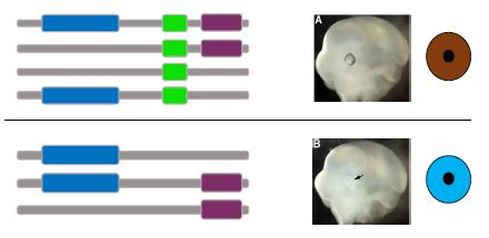
The purpose of aim 1 was to discover the domain(s) of MITF that is responsible for DNA binding and transcriptional activation of the tyrosinase promoter. First, SMART database is used to discover what domains the MITF protein has. Then protein constructs will be made that include or exclude only one domain. These will then be put into mice. I will then look for the mice with brown eyes, as this means proper binding of the MITF protein to the tyrosinase promoter. I hypothesize that the constructs with the HLH domain will result in wildtype brown eyed mice.
Then to further test the HLH domain for specific amino acids that are important for binding, I will create random mutations in the HLH domain using CRISPR/Cas9. Then the mice with incomplete penetrance will be assayed using an electrophoretic mobility shift assay (EMSA), to test for the binding affinity of the mutant MITF proteins to the tyrosinase promoter. I suspect to find that these mice will have some binding and some non-bound DNA.
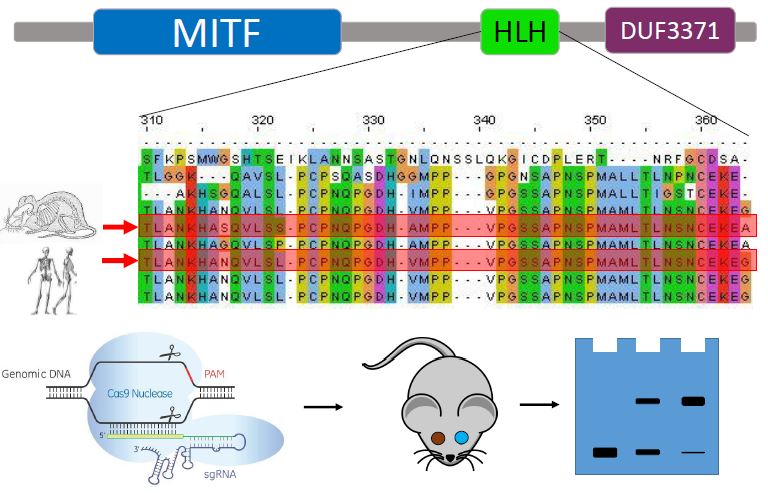
Figure 2. (A) Clustal Omega alignment of the HLH domain across species. Random mutations will be made within the HLH domain. (B) Procedure outline. CRISPR/Cas9 will create mutations, and mice with incomplete penetrance will be assayed. In the gel the middle column shows weak binding of the MITF protein to the tyrosinase promoter. [4-6]
Aim 2
|
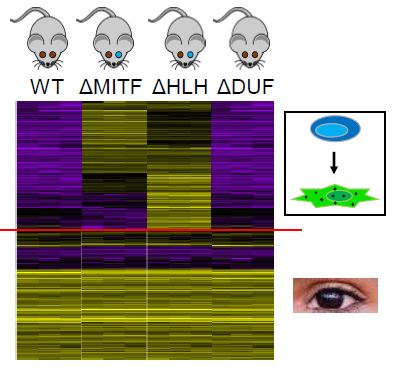
The purpose of aim 2 is to understand if gene expression of melanocyte genes or eye development genes would be affected by MITF mutations. Using RNA-seq on WT, MITF knockout, HLH domain knockout, and DUF domain knockout mice. WT and DUF mice are used as the controls. I suspect that in the MITF knockout and HLH domain knockout, gene expression of melanocyte genes will be affected because there is no longer melanocyte differentiation, but gene expression of eye development genes will not be affected because the eyes are still going to develop.
|
Aim 3
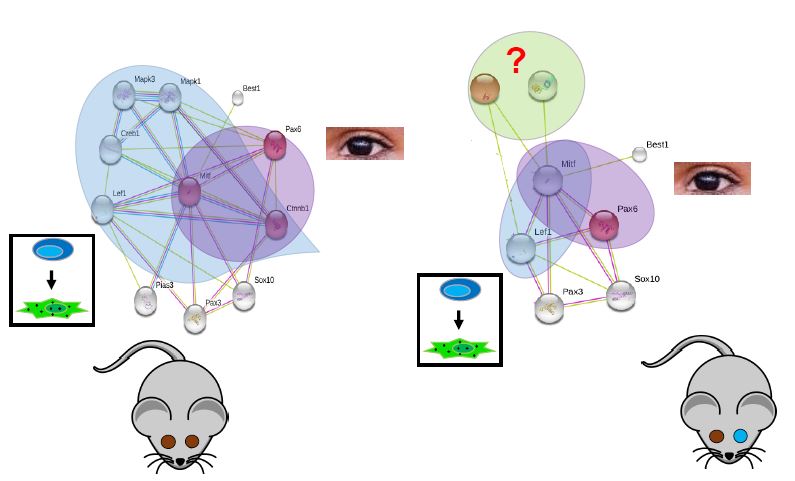
The purpose of aim 3 is to discover which protein interactions are different in WT vs WS2A individuals. After using STRING to map WT protein interactions, incomplete penetrance mice, with one brown and one blue eye, will be assayed using Co-Immunoprecipitation, followed by liquid chromatography separation and tandem mass spec. This will show what protein interactions are taking place within mutant MITF individuals.
I hypothesize that there will be a decrease in MITFs interactions with the melanocyte differentiation proteins, as melanocyte differentiation will no longer be happening completely. There is also the possibility that new interactions will be formed when MITF in mutated. If this is the case, the new proteins can be validated by creating knockouts mice and screening for a change in phenotype. If they do play a role, we would see incomplete or complete loss of pigmentation in the eyes.
Future Directions
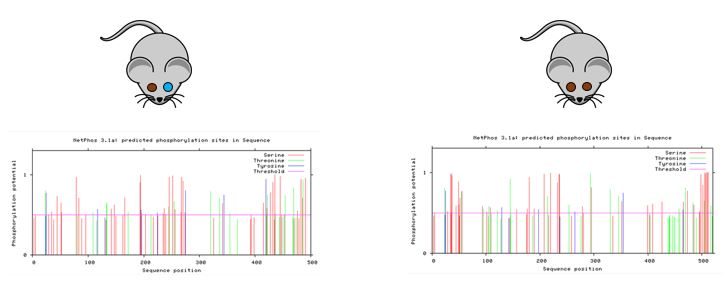
I think the next step would be to take the incomplete penetrance mutants and compare phosphorylation of the MITF gene to the WT mice. For activation of MITF it becomes phosphorylated. Comparing these sites may give insight to what is happening when MITF fails to activate the tyrosinase promoter and induce melanocyte differentiation.
References
1. Shi, Y., Li, X., Ju, D., Li, Y., Zhang, X., & Zhang, Y. (2016). A novel mutation of the MITF gene in a family with Waardenburg syndrome type2: A case report. Experimental and Therapeutic Medicine. doi:10.3892/etm.2016.3042
2. Tachibana, M. (1997). Evidence to Suggest That Expression of MITF Induces Melanocyte Differentiation and Haploinsufficiency of MITF Causes Waardenburg Syndrome Type 2A. Pigment Cell Research,10(1-2), 25-33. doi:10.1111/j.1600-0749.1997.tb00462.x
3. Westenskow, P., Piccolo, S., & Fuhrmann, S. (2009). -catenin controls differentiation of the retinal pigment epithelium in the mouse optic cup by regulating Mitf and Otx2 expression. Development,136(15), 2505-2510. doi:10.1242/dev.032136
4. CRISPR Guide RNA. (n.d.). Retrieved May 07, 2017, from http://dharmacon.gelifesciences.com/gene-editing/crispr-cas9/crispr-guide-rna/
5. Simple Cartoon Mouse clip art. (n.d.). Retrieved May 07, 2017, from http://www.clker.com/clipart-simple-cartoon-mouse-2.html
6. http://www.ebi.ac.uk/Tools/msa/clustalo/
7. http://shahidulsahel.com/travel-and-street/
8. https://github.com/trinityrnaseq/trinityrnaseq/wiki/Trinity-Differential-Expression
9. http://string-db.org/
10. http://www.cbs.dtu.dk/services/NetPhos/
1. Shi, Y., Li, X., Ju, D., Li, Y., Zhang, X., & Zhang, Y. (2016). A novel mutation of the MITF gene in a family with Waardenburg syndrome type2: A case report. Experimental and Therapeutic Medicine. doi:10.3892/etm.2016.3042
2. Tachibana, M. (1997). Evidence to Suggest That Expression of MITF Induces Melanocyte Differentiation and Haploinsufficiency of MITF Causes Waardenburg Syndrome Type 2A. Pigment Cell Research,10(1-2), 25-33. doi:10.1111/j.1600-0749.1997.tb00462.x
3. Westenskow, P., Piccolo, S., & Fuhrmann, S. (2009). -catenin controls differentiation of the retinal pigment epithelium in the mouse optic cup by regulating Mitf and Otx2 expression. Development,136(15), 2505-2510. doi:10.1242/dev.032136
4. CRISPR Guide RNA. (n.d.). Retrieved May 07, 2017, from http://dharmacon.gelifesciences.com/gene-editing/crispr-cas9/crispr-guide-rna/
5. Simple Cartoon Mouse clip art. (n.d.). Retrieved May 07, 2017, from http://www.clker.com/clipart-simple-cartoon-mouse-2.html
6. http://www.ebi.ac.uk/Tools/msa/clustalo/
7. http://shahidulsahel.com/travel-and-street/
8. https://github.com/trinityrnaseq/trinityrnaseq/wiki/Trinity-Differential-Expression
9. http://string-db.org/
10. http://www.cbs.dtu.dk/services/NetPhos/
Final Presentation
| baars_final_presentation_4.25.2017.pdf | |
| File Size: | 1698 kb |
| File Type: | |
Draft 1
| baars_presentation_draft_1_2.21.17.pptx | |
| File Size: | 1143 kb |
| File Type: | pptx |
| jboles.comments_baars_presentation_draft_1_2.21.17.pptx | |
| File Size: | 1327 kb |
| File Type: | pptx |
Draft 2
| baars_presentation_draft_4.5.2017.pptx | |
| File Size: | 3361 kb |
| File Type: | pptx |
| baars_presentation_draft_4.5.2017.pptx | |
| File Size: | 3361 kb |
| File Type: | pptx |